Chinese Journal of Tissue Engineering Research ›› 2019, Vol. 23 ›› Issue (19): 2980-2985.doi: 10.3969/j.issn.2095-4344.1243
Previous Articles Next Articles
Association between single nucleotide polymorphism (8q24.21) and lumbar degeneration disease in 800 cases from Guangxi Zhuang Autonomous Region
Liu Yang, Yang Qinghua, Guan Yewen, Wang Jiaqi, Jiang Hua
- (Department of Spinal Surgery, the First Affiliated Hospital of Guangxi Medical University, Nanning 530021, Guangxi Zhuang Autonomous Region, China)
-
Received:2019-02-19Online:2019-07-08Published:2019-07-08 -
Contact:Jiang Hua, MD, Associate chief physician, Master’s supervisor, Department of Spinal Surgery, the First Affiliated Hospital of Guangxi Medical University, Nanning 530021, Guangxi Zhuang Autonomous Region, China -
About author:Liu Yang, Master candidate, Department of Spinal Surgery, the First Affiliated Hospital of Guangxi Medical University, Nanning 530021, Guangxi Zhuang Autonomous Region, China -
Supported by:the National Natural Science Foundation of China, No. 81460353 and 81860406 (both to JH); the Natural Science Foundation of Guangxi Zhuang Autonomous Region, No. 2015GXNSFBA139167 (to JH)
CLC Number:
Cite this article
Liu Yang, Yang Qinghua, Guan Yewen, Wang Jiaqi, Jiang Hua. Association between single nucleotide polymorphism (8q24.21) and lumbar degeneration disease in 800 cases from Guangxi Zhuang Autonomous Region [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(19): 2980-2985.
share this article
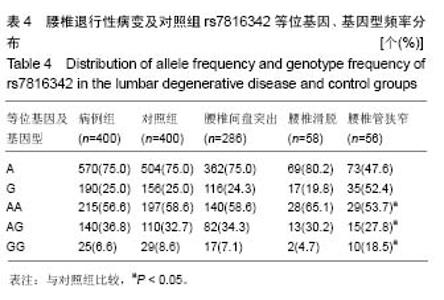
2.2 8q24.21区域rs7816342、rs4130415位点的检测结果 ABI7500型实时荧光定量PCR系统测得的SNP结果显示,rs7816342位点的基因型有3种,分别是AA、AG、GG;rs4130415位点的基因型有3种,分别是TT、TC、CC。 2.3 Hardy-Weinberg平衡定律检测 经χ2检验,病例组和对照组的研究对象在该群体均存在Hardy-Weinberg平衡(P > 0.05)。 2.4 8q24.21区域rs7816342、rs4130415位点与腰椎退行性疾病的相关性关系 2.4.1 rs7816342位点在病例组及对照组中的各等位基因、基因型频率 见表4,A和G等位基因频率在病例组分别为75.0%和25.0%,在对照组分别占75.0%和25.0%,2组间等位基因频率分布差异无显著性意义(χ2=0.00,P=0.999,OR=1.000,95%CI:0.787-1.271),2组间的基因型频率分布差异无显著性意义(χ2=1.986,P=0.370)。 在亚组分析中,腰椎间盘突出组(亚组1)的A和G等位基因频率在腰椎退行性疾病中分别为75.7%、24.3%,在对照组中分别占75.0%、25.0%,2组间等位基因频率分布差异无显著性意义(χ2=0.085,P=0.771,OR=0.961,95%CI=0.737-1.254),2组间的基因型频率分布差异无显著性意义(χ2=0.426,P=0.808)。 腰椎滑脱组(亚组2)组的A和G等位基因频率在腰椎退行性疾病中分别为80.2%、19.8%,在对照组中分别占75.0%、25.0%,2组间等位基因频率分布差异无显著性意义(χ2=1.145,P=0.285,OR=0.739,95%CI=0.424- 1.288),2组间的基因型频率分布差异无显著性意义(χ2= 1.182,P=0.554)。"
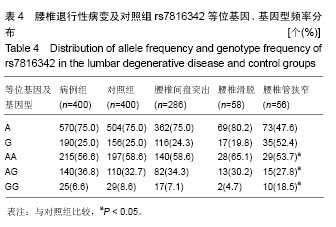
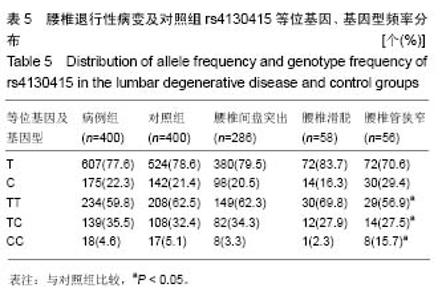
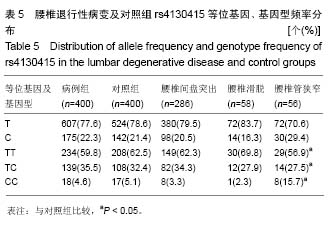
腰椎管狭窄组(亚组3)和对照组的A和G等位基因频率在腰椎退行性疾病中分别为:47.6%、52.4%,在对照组中分别占75.0%、25.0%,2组间等位基因分布差异无显著性意义(χ2=2.702,P=0.100,OR=1.438,95%CI= 0.931-2.222),2 组间的基因型频率分布差异有显著性意义(χ2=9.515,P=0.008)。 2.4.2 rs4130415位点在病例组及对照组中的各等位基因、基因型频率 见表5,T和C等位基因频率在病例组中分别为78.6%、21.4%,在对照组中分别占77.6%、22.3%,2组间等位基因频率分布差异无显著性意义(χ2=0.235,P=0.628,OR=0.941,95%CI=0.782-1.207),2组间的基因型频率分布差异无显著性意义(χ2=0.807,P=0.668)。"
| [1] Melrose J. Strategies in regenerative medicine for intervertebral disc repair using mesenchymal stem cells and bioscaffolds. Regen Med. 2016;11(7):705-724.[2] Stubbs B,Koyanagi A,Thompson T,et al. The epidemiology of back pain and its relationship with depression, psychosis, anxiety, sleep disturbances, and stress sensitivity: Data from 43 low- and middle-income countries. Gen Hosp Psychiatry.2016;43:63-70.[3] Shmagel A, Foley Rand Ibrahim H. Epidemiology of Chronic Low Back Pain in US Adults: Data From the 2009-2010 National Health and Nutrition Examination Survey. Arthritis Care Res (Hoboken). 2016;68(11):1688-1694.[4] Cheung KM,Karppinen J,Chan D, et al.Prevalence and pattern of lumbar magnetic resonance imaging changes in a population study of one thousand forty-three individuals. Spine (Phila Pa 1976). 2009;34(9):934-940.[5] Takatalo J,Karppinen J,Niinimaki J,et al. Does Lumbar Disc Degeneration on Magnetic Resonance Imaging Associate With Low Back Symptom Severity in Young Finnish Adults?.Spine.2011; 36(25):2180-2189.[6] Dario AB,Ferreira ML,Refshauge KM,et al.The relationship between obesity, low back pain, and lumbar disc degeneration when genetics and the environment are considered: a systematic review of twin studies.Spine J.2015;15(5):1106-1117.[7] Battie MC,Videman T. Lumbar disc degeneration: Epidemiology and genetics. J Bone Joint Surg Am. 2006;88 Suppl 2:3-9. [8] Tarnoki AD,Tarnoki DL,Kovacs DT,et al.Genetic relationship between obstructive sleep apnea and lumbar disc degeneration: results from a twin study. European Respiratory Journal. 2017; 50.[9] Zhou X,Cheung CL,Karasugi T,et al.Trans-Ethnic Polygenic Analysis Supports Genetic Overlaps of Lumbar Disc Degeneration With Height, Body Mass Index, and Bone Mineral Density. Front Genet. 2018;9:267.[10] Battie MC, Videman Tand Parent E. Lumbar disc degeneration - Epidemiology and genetic influences. Spine.2004;29(23): 2679-2690.[11] Bjornsdottir G, Benonisdottir S, Sveinbjornsson G, et al. Sequence variant at 8q24.21 associates with sciatica caused by lumbar disc herniation. Nature Communications.2017, 8.[12] Baldwin Aand Mesfin A. Adjacent Segment Disease 44 Years Following Posterior Spinal Fusion for Congenital Lumbar Kyphosis. Spine Deform.2017;5(6):435-439.[13] Dong L,Xu Z,Chen X,et al.The change of adjacent segment after cervical disc arthroplasty compared with anterior cervical discectomy and fusion: a meta-analysis of randomized controlled trials. Spine J.2017;17(10):1549-1558.[14] Arnbak B,Jensen RK,Manniche C,et al.Identification of subgroups of inflammatory and degenerative MRI findings in the spine and sacroiliac joints: a latent class analysis of 1037 patients with persistent low back pain.Arthritis Res Ther.2016;18(1):237.[15] Muriuki MG, Havey RM,Voronov LI,et al. Effects of motion segment level, Pfirrmann intervertebral disc degeneration grade and gender on lumbar spine kinematics.J Orthop Res.2016; 34(8):1389-1398.[16] Xiao L,Ni CL,Shi JD,et al.Analysis of Correlation Between Vertebral Endplate Change and Lumbar Disc Degeneration. Medical Science Monitor.2017;23:4932-4938.[17] Wu B,Meng C,Wang H,et al.Changes of proteoglycan and collagen II of the adjacent intervertebral disc in the cervical instability models. Biomed Pharmacother.2016; 84:754-758.[18] Grant MP,Epure LM,Bokhari R,et al.Human cartilaginous endplate degeneration is induced by calcium and the extracellular calcium-sensing receptor in the intervertebral disc. Eur Cell Mater. 2016;32:137-151.[19] Xu HG,Zheng Q,Song JX,et al.Intermittent cyclic mechanical tension promotes endplate cartilage degeneration via canonical Wnt signaling pathway and E-cadherin/beta-catenin complex cross-talk. Osteoarthritis Cartilage. 2016;24(1):158-168.[20] Sivan SS, Hayes AJ, Wachtel E, et al. Biochemical composition and turnover of the extracellular matrix of the normal and degenerate intervertebral disc. Eur Spine J.2014;23 Suppl 3: S344-53.[21] Garnero P. The Role of Collagen Organization on the Properties of Bone. Calcif Tissue Int. 2015,97(3):229-240.[22] 张学利,宋海峰,朱如森,等.Ⅸ型胶原蛋白A2基因多态性与环境因素在腰椎间盘退变疾病中的作用[J] 中华实验外科杂志,2017, 34(7): 1232-1235.[23] Wu HH, Wang ST, Chen WY, et al. Collagen IX gene polymorphisms and lumbar disc degeneration: a systematic review and meta-analysis. J Orthop Surg Res. 2018;13(1):47.[24] Mio F, Chiba K, Hirose Y, et al. A functional polymorphism in COL11A1, which encodes the alpha 1 chain of type XI collagen, is associated with susceptibility to lumbar disc herniation[J]. Am J Hum Genet.2007;81(6):1271-1277.[25] Ruel N, Markova DZ,Adams SL,et al.Fibronectin fragments and the cleaving enzyme ADAM-8 in the degenerative human intervertebral disc.Spine (Phila Pa 1976).2014; 39(16):1274-1279.[26] Urano T,Narusawa K,Shiraki M,et al.Single-nucleotide polymorphism in the hyaluronan and proteoglycan link protein 1 (HAPLN1) gene is associated with spinal osteophyte formation and disc degeneration in Japanese women. Eur Spine J.2011; 20(4):572-577.[27] S M H, M K S, G K C, et al. Association of CILP, COL9A2 and MMP3 Gene Polymorphisms with Lumbar Disc Degeneration in an Indian Population.J Mol Neurosci.2018;66(3):378-382.[28] Li Y, Li K, Han XG, et al. The imbalance between TIMP3 and matrix-degrading enzymes plays an important role in intervertebral disc degeneration. Biochemical and Biophysical Research Communications, 2016, 469(3):507-514.[29] 张博,杨少华,林世洲,等.基质金属蛋白酶1在不同程度退变腰椎间盘髓核与纤维环中的表达及意义 [J].中国组织工程研究, 2012,16(13): 2346-2349.[30] Deng B, Ren JZ, Meng XQ, et al. Expression profiles of MMP-1 and TIMP-1 in lumbar intervertebral disc degeneration. Genet Mol Res.2015;14(4):19080-19086.[31] Yuan HY, Tang Y, Lei L, et al.[Synergistic interaction between MMP-3, VDR gene polymorphisms and occupational risk factors on lumbar disc degeneration]. Zhonghua Lao Dong Wei Sheng Zhi Ye Bing Za Zhi.2010;28(5):334-338.[32] Williams FMK, Bansal AT, van Meurs JB, et al. Novel genetic variants associated with lumbar disc degeneration in northern Europeans: a meta-analysis of 4600 subjects. Annals of the Rheumatic Diseases, 2013, 72(7):1141-1148.[33] Lin WP,Lin JH,Chen XW,et al.Interleukin-10 promoter polymorphisms associated with susceptibility to lumbar disc degeneration in a Chinese cohort.Genet Mol Res.2011;10(3): 1719-1727.[34] Omair A,Lie BA,Reikeras O, et al. An Association Study of Interleukin 18 Receptor Genes (IL18R1 and IL18RAP) in Lumbar Disc Degeneration. Open Orthop J.2012; 6:164-171.[35] Jiang H, Qin Z,Zong S,et al. Vitamin D receptor gene polymorphisms and lumbar disc degeneration: a systematic review and meta-analysis. Eur Spine J.2017;26(1):267-277.[36] Zawilla NH, Darweesh H, Mansour N,et al. Matrix metalloproteinase-3, vitamin D receptor gene polymorphisms, and occupational risk factors in lumbar disc degeneration. J Occup Rehabil.2014; 24(2):370-381.[37] Yuan HY,Tang Y,Liang YX,et al.Matrix metalloproteinase-3 and vitamin d receptor genetic polymorphisms, and their interactions with occupational exposure in lumbar disc degeneration. J Occup Health.2010;52(1):23-30.[38] Schultz DR,Harrington WJ Jr.. Apoptosis: Programmed cell death at a molecular level. Semin Arthritis Rheum. 2003;32(6):345-369.[39] Ariga K,Yonenobu K,Nakase T,et al.Mechanical stress-induced apoptosis of endplate chondrocytes in organ-cultured mouse intervertebral discs - An ex vivo study.Spine.2003; 28(14): 1528-1533.[40] Sakai D, Nakamura Y, Nakai T, et al. Exhaustion of nucleus pulposus progenitor cells with ageing and degeneration of the intervertebral disc. Nat Commun.2012;3:1264.[41] Jiang M, Zhu K, Grenet J, et al.Retinoic acid induces caspase-8 transcription via phospho-CREB and increases apoptotic responses to death stimuli in neuroblastoma cells. Biochim Biophys Acta. 2008;1783(6):1055-1067.[42] Wang X, Luo T, Ruan M, et al. Association of the CCDC26 rs4295627 polymorphism with the risk of glioma: Evidence from 7,290 cases and 11,630 controls. Mol Clin Oncol.2016;4(5): 878-882.[43] Cao K, Li M, Miao J, et al. CCDC26 knockdown enhances resistance of gastrointestinal stromal tumor cells to imatinib by interacting with c-KIT.Am J Transl Res.2018;10(1):274-282.[44] Zeng J, Luo Y, Yu M, et al.CCDC26 rs4295627 polymorphisms associated with an increased risk of glioma: A meta-analysis. Adv Clin Exp Med.2017; 26(8):1275-1281.[45] Peng Wand Jiang A.Long noncoding RNA CCDC26 as a potential predictor biomarker contributes to tumorigenesis in pancreatic cancer. Biomed Pharmacother.2016;83:712-717.[46] Wang SL, Hui YZ, Li XM, et al. Silencing of lncRNA CCDC26 Restrains the Growth and Migration of Glioma Cells In Vitro and In Vivo via Targeting miR-203.Oncology Res.2018; 26(8):1143-1154.[47] Izadifard M, Pashaiefar H, Yaghmaie M, et al. Expression Analysis of PVT1, CCDC26, and CCAT1 Long Noncoding RNAs in Acute Myeloid Leukemia Patients. Genet Test Mol Biomarkers. 2018;22(10):593-598. [48] Vaubel R,Kollmeyer T,Caron A,et al.Synchronous gemistocytic astrocytoma IDH-mutant and oligodendroglioma IDH-mutant and 1p19q-codeleted in a patient with CCDC26 polymorphism. Acta Neuropathol. 2017;134(2):317-319. |
| [1] | He Liumei, Chen Hao, Zhong Yanping, Quan Zhanrou, Zou Hongyan. Next-generation sequencing of identifying the human leukocyte antigen-A*24:353 and its gene frequency analysis [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(25): 4009-4012. |
| [2] | Gao Ziqing, Ruan Sibei, Li Li, Ling Feng, Tang Xiaoqin, Kang Qingmei, Luo Siyi, Luo Jing, Tang Yaping, Tang Mingxi . Identification and survival analysis of tTA/tetO-CCKR-2 double transgenic mice [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(11): 1682-1687. |
| [3] | Yang Jinfeng, Ma Sanhui. Association of HTRA1 and HAPLN1 gene polymorphism with intervertebral disc degeneration [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(32): 5128-5132. |
| [4] | Zhou Wenming, Lin Yifeng, Zhang Zhen, Chi Liye. Effect of Bushen Zhuangdu Fang serum on mitochondrial apoptotic pathway of nucleus pulposus cells [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(23): 3643-3648. |
| [5] | Zhong Yanping, Zou Hongyan, Quan Zhanrou, Deng Zhihui, Hong Wenxu. Analysis of full-length sequence and 18 point mutations of HLA-B in a leukemia patient and her family [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(1): 77-82. |
| [6] | Huang Qun, Sheng Xiaolei, Yan Fei, Zhou Zhiping, Zhu Xianwei. Minimally invasive transforaminal lumbar interbody fusion through Quadrant channel for treatment of lumbar degenerative diseases [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(36): 5818-5823. |
| [7] | Lu Qiuping1, Gui Yuchang2, Xu Jianwen2, Qin Haibiao3, Rao Yuansen1. Association between vitamin D receptor gene polymorphism and semi-quantitative classification of thoracolumbar osteoporotic vertebral compression fractures [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(35): 5577-5581. |
| [8] | Quan Zhanrou, He Liumei, Chen Hao, Hong Wenxu, Gao Suqing . Distribution characteristics of polymorphism of human leukocyte antigen C of hepatitis B virus carriers in patients from Shenzhen [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(3): 482-486. |
| [9] | Xue Zhixing, Zhou Jianwei, Chi Cheng, Wang Fei, Ma Yuquan. Long-term effects of inorganic osteogenesis-inducing scaffold versus autologous bone in lumbar interbody fusion: protocol for a non-randomized, controlled trial with 2-year follow-up [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(22): 3518-3521. |
| [10] | Li Pengfei, Wang Tao, Ma Xinlong . Association between COL9A2 gene polymorphisms and intervertebral disc degeneration in Asian: a meta-analysis [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(20): 3275-3280. |
| [11] | Long Yanming, Xie Mengsheng, Xue Wenli, Huang Jiajie, Li Xiaojie. Characteristics of bone marrow mesenchymal stem cells in a rat model of retinoic acid-induced osteoporosis [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(17): 2637-2643. |
| [12] | Sun Ke, Yang Xuejun. Application of finite element technology in lumbar degenerative diseases [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(16): 2553-2559. |
| [13] | He Liu-mei, Wang Song-xing, Xu Yun-ping. Identification of genomic full-length sequence of human leukocyte antigen-E and its two novel alleles [J]. Chinese Journal of Tissue Engineering Research, 2017, 21(25): 4068-4074. |
| [14] | Liu Xiao-hua, Chi Xiao-yun, Jiao Shu-xian. Identification of novel HLA alleles: HLA-A*24:233 and HLA-A*26:89 [J]. Chinese Journal of Tissue Engineering Research, 2015, 19(6): 950-954. |
| [15] | Xu Rui, Yang Yi-ning, Ma Yi-tong, Li Xiao-mei, Zhao Qian, Chen Bang-dang, Liu Fen . Association between the rs1007888 polymorphism of macrophage migration inhibitory factor gene and coronary heart disease in the Kazakhs of China [J]. Chinese Journal of Tissue Engineering Research, 2015, 19(2): 231-235. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||